Research
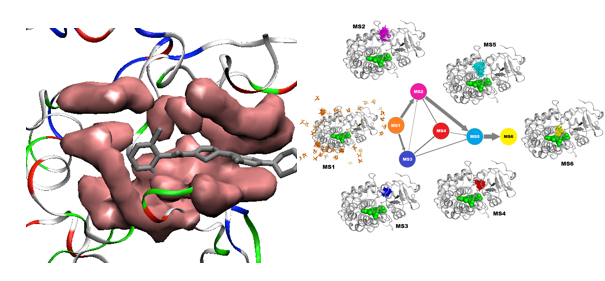
Simulation of Protein-ligand recognition kinetics
-
Classical Molecular Dynamics Simulation
-
Atomistic and Physics-based modelling
-
Illustration of multiscale mechanism
-
Statistical Models and machine learning
-
Intensive collaboration with experiments
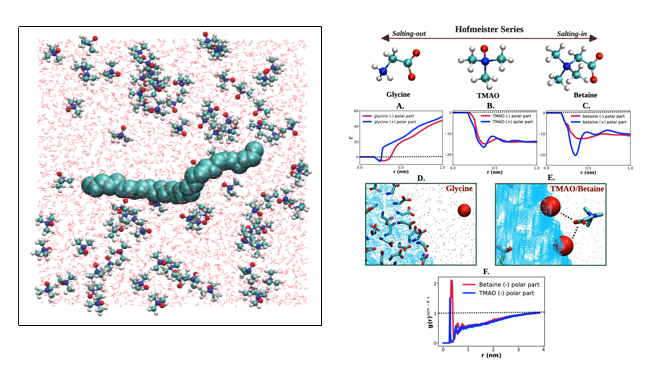
Macromolecule/Interface/Osmolyte
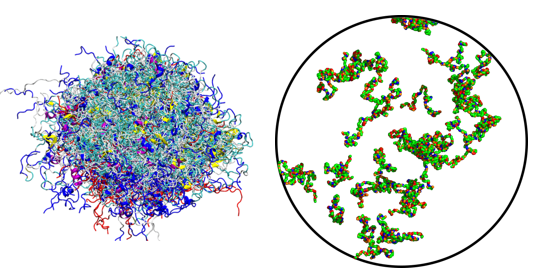
Conformation & Phase separation Of Intrinsically Disordered protein
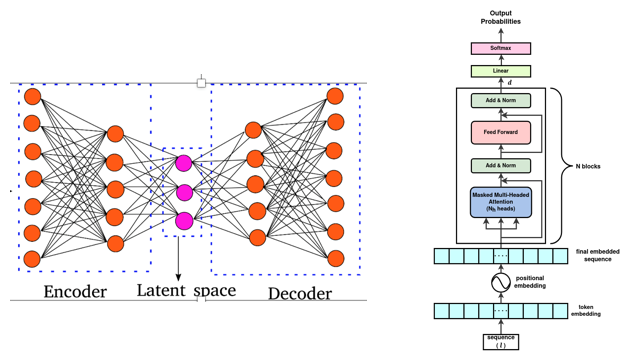
Machine-Learning and generative AI
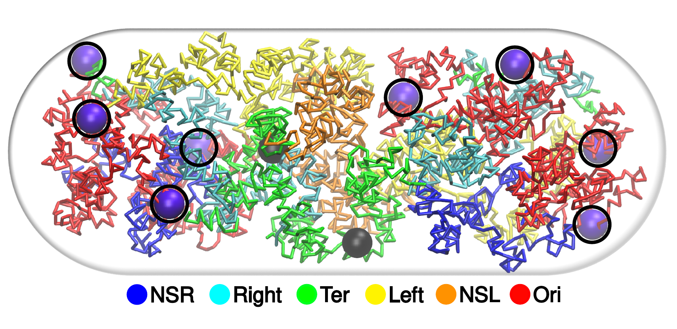
Integrative modelling of bacterial chromosome